REEU Projects
**Applications for the 2026 program are closed*
2026 REEU Research Projects
You do not need to already have the skills mentioned in the project descriptions, just an interest to learn. You will gain hands-on training during the program.
List three projects on your application. We cannot guarantee that you will be placed in one of your top three selections, but your selections will help us place you in a project that is a good match for your interests.
Note - Projects are subject to change if research priorities evolve in the mentor lab before the start of the program.
Broadening phenotypic diversity of wheat by high-throughput genome editingCRISPR/CAS9 gene editing has emerged as a disruptive technology that can take full advantage of the available genomic information and help to rapidly assess the role of newly identified genes in controlling agronomic traits, generate new variants of genes to broaden genetic diversity available for breeding, and to transfer this newly created variation into wheat varieties. This technology relies on the ability of the CAS proteins directed by a single-guide RNA (sgRNA) to introduce changes into DNA at a specific location in genome, usually within the genes. The utility of the CRISPR-CAS system for gene editing (GE) was successfully demonstrated in various crops including wheat. Currently, our group applies the high-throughput CAS technology to edit genes for improving yield, nutritional value, and disease resistance in wheat. Typically, gene editing project involves selection of a gene target that can affect positively the trait of interest. Then, a set of gRNAs targeting this gene is designed using bioinformatical tools followed by validation of these gRNAs in the wheat protoplasts, a specially prepared cells with the cell wall removed. The efficiency of editing of gene targets in the wheat protoplasts is tested by next-generation sequencing (NGS) at the Integrated Genomics Facility. The intern will conduct these experiments and bioinformatical analysis of NGS data as part of the gene editing projects aimed at improving wheat traits. 60% Lab, 40% Computer Mentors: Eduard Akhunov |
Quantifying the inputs and costs required to establish and maintain seeded turfgrass compared to sodded turfgrassThis field research will assess and quantify plant growth, development, and associated inputs and expenses needed to establish and maintain seeded vs. sodded turf of multiple species. The undergraduate student will observe and quantify the effects of environmental conditions, time of year, and different management practices on turfgrass systems. The undergraduate student will use digital data collection and analysis tools in the field. This research will help educate turfgrass and landscape practitioners on the differences in establishment and management costs, practices, and challenges associated with establishment methods and specific species that will reduce management inputs in turfgrass systems. This project will have a duration of 9-weeks, interacting with a research team and allowing participation in the other research activities, professional development, field trips, and sessions. My team's grad students and post-docs will help with mentoring. I will participate in the mentor training sessions offered each year before the interns’ arrival and communicate effectively with my intern before arrival to increase readiness and help ensure they maximize their time in the program. I am committed to assisting in the longer-term development of the REEU participants by providing guidance and letters of reference after the internship. I will include opportunities for undergraduate internships in future grant proposals where it is suitable and permitted by the funding agency. Mentor: Ross Braun |
Putting it all together- Long-read sequencing and assembling a fungal genome
Areas: Genomics, mycology, molecular biology, bioinformatics and plant pathology Mentor: David Cook |
Stopping a Cereal Killer!Wheat blast, caused by the fungal pathogen Magnaporthe oryzae Triticum (MoT), is a devastating emerging disease historically restricted to South America, but more recently spread to southern Asia and Africa. Genetic resistance is the most sustainable mitigation strategy, but its sources are rare. The main currently deployed resistance is only partially effective and is being overcome by newly emerged, highly aggressive strains. M. oryzae infects more than 50 grass species that may contribute alternate sources of inoculum and facilitate genetic recombination and variation, thus posing an immediate risk of a host jump and dangerous new pathogen populations. You will learn different laboratory techniques related to genetic resistance and fungal and plant biology such as pathogen isolation, DNA extraction, microscopy, inoculation, and disease evaluation, among others. Additionally, you will be able to visit the BRI and observe research conducted under BSL-3 conditions. The research will comprise roughly 80% lab, 10% computer, and 10% greenhouse work. Mentor: Giovana Cruppe |
Synthetic Biology to Make Super SeedsAcetyl-triacylglycerols (acetyl-TAG) are triacylglycerols that possess an sn-3 acetate group instead of a long chain fatty acid. This unusual structure confers useful properties to acetyl-TAG, including reduced kinematic viscosity and improved cold temperature performance. Acetyl-TAG can by synthesized in transgenic oilseeds by expressing a diacyglycerol acetyltransferase enzyme that was isolated from plant species that naturally produce acetyl-TAG. The levels of acetyl-TAG produced in transgenic seeds can be further enhanced by manipulating endogenous metabolic pathways, for example by eliminating competing reactions using RNA interference or CRISPR-based genome editing. Interns will gain experience creating plant transformation vectors to alter endogenous metabolism, generating and characterizing transgenic plants, and different lipid analysis techniques, such as gas chromatography or electro-spray ionization mass spectrometry (ESI-MS). Mentor: Tim Durrett |
Microbial inoculants for biocontrolThe Hansen lab has recently used its patented microwell technology platform to screen rhizobacteria culture collections for non-pathogenic bacterial isolates that antagonize the plant pathogen Agrobacterium tumefaciens during co-culture. A. tumefaciens is the agent of Crown Gall Disease, affecting more than 90 families of dicotyledonous plants and resulting in significant agronomic losses. While our panel of isolates has potential to inhibit A. tumefaciens, investigation of these isolates and their interactions with A. tumefaciens in the context of a plant root host and in various soil environments is required. The REEU student will investigate the ability of our selected rhizobacteria isolates to protect against Crown Gall Disease by inhibiting A. tumefaciens colonization into the roots of developing Helianthus annuus (sunflower) plants, first in lab-scale plant culture studies, and then within the soil in greenhouse environments. This work will progress the discovery and application of robust biocontrol agents for pathogen protection, alleviating the detrimental environmental and health effects associated with synthetic agrochemicals. Mentor: Ryan Hansen |
Let’s Make Better WheatWe will screen wheat breeding populations for pathogen and disease resistance in natural field conditions to identify the genetic factors that help produce quality breads and cereals. The intern will rate wheat’s response to several pathogens, and using high throughput sequencing and genotyping techniques identify genetic regions involved in plant resistance using bioinformatics tools. These novel genetic factors will be transferred into our breeding programs for improved wheat for future cultivar development and deployment. Mentor: Katherine Jordan |
Wild Wheat!
Areas: genetics and genomics, plant breeding, plant pathology, cytogenetics. Greenhouse 25%, Lab 75% Mentor: Dal-Hoe Koo |
Weed Management
Field 80%, Lab + Greenhouse 20% Mentor: Sarah Lancaster |
Genome mapping of plant pathogens
Lab 40%, Computational work 60% Mentor: Sanzhen Liu |
Molecular Switches in Plant ImmunityPlants cannot run away from pathogens; instead, they rapidly activate powerful immune responses. Within minutes of detecting infection, plant cells start to reprogram the entire cell to defend themselves. How can this happen so quickly? The answer lies in post-translational modifications (PTMs) — small chemical changes such as phosphorylation or ubiquitination that fine-tune proteins after they are made. Rather than creating new proteins from scratch, cells quickly adjust the activity, stability, or interactions of existing proteins, enabling a rapid and precise defense response. Just like Optimus Prime shifting from truck mode to robot mode, these modifications can dramatically alter what a protein does without changing what it fundamentally is. Our lab studies how these molecular switches control plant immunity at the biochemical and molecular level. In this project, you will clone a gene into a plasmid, express and purify a plant immune protein, analyze its post-translational modifications, and test how these modifications regulate its activity. You will gain hands-on experience in molecular cloning, protein biochemistry, and plant biology while discovering how plants defend themselves at the molecular level. Mentor: Xiyu Ma |
Blending biology with technology to manage arthropods in field cropsKansas crop growers face a diverse array of insect pests. Consequently, digital entomology plays a crucial role in addressing complex problems in field crops by using innovative technologies, such as remote sensing, data analytics, and machine learning to monitor insect populations and their behavior. This approach enables early detection of pest outbreaks, precise intervention strategies, and sustainable pest management practices, ultimately safeguarding crop yields and food security. In this project, you will learn how to use cutting edge technology like deep learning to help develop integrated pest management methods and help develop and deliver information to growers using novel mobile technologies. You will learn to scout and sample insects in the fields, map their prevalence, develop sophisticated sampling tools using computer vision, and contribute to Extension programs. Lab 25%/Field 50%/Computer 25% Mentor: Brian McCornack |
Exploring wheat wild relatives for virus tolerance and resistance and the molecular mechanism of virus tolerance
My research aims to explore viral tolerance as a breeding tool. From an evolutionary perspective, tolerance is an evolutionary mechanism for both viruses and their hosts for adaptation. Using tolerance instead of resistance in the field to impose less selection pressure on plant viruses has recently received attention. However, the molecular mechanisms of tolerance are not clear. My research aims to address these knowledge gaps using wheat wild relatives, Aegilops tauschii genotypes, and two important wheat viruses: WSMV and TriMV. Our hypothesis is that the virus not only changes the gene expression profile of the host, but also the host epigenetics (non-coding RNAs including miRNAs pattern) as well as proteomic and metabolic profiles. Mentor: Shahideh Nouri |
Characterizing a newly discovered virus associated with wheatMy lab recently discovered a novel virus in KS wheat fields. This emerging virus named Wheat umbra-like virus (WULV) belongs to a new group of subviral RNAs recently identified in several plant species. To have a better understanding of the biology of WULV and its potential as new pathogens of wheat in the region, we aim to characterize it using traditional and molecular virology and genomics. Mentor: Shahideh Nouri |
Engaging horticulture and agronomic professionals through extension and plant diagnosticsFrom agronomic crops to fruit, vegetables, trees, and turf, the plant sciences industry is diverse! With so many plant species, there are many diseases and environmental stresses for professionals and home gardeners to understand. In this extension-focused internship, there will be an opportunity to learn plant disease diagnostic techniques, develop engaging online educational materials (video and written), assist with developing and implementing hands-on educational activities for broad audiences, learn about program development and evaluation, spend time “shadowing” county agents, and conduct outreach to K-12 students and teachers. Learn plant pathology by teaching it to others! Mentors: Judy O'Mara and Megan Kennelly |
Insect hormone disruptors
Mentor: Yoonseong Park |
My Feet Hurt!Over the last eight years, the Applied Wheat Pathology lab has developed and maintained a robust field experiment exploring the effect of multiple root rot pathogens on wheat production. We recently began incorporating perennial plants under development by The Land Institute to study their ability to withstand a pathogen onslaught in the rhizosphere. The student working on this project will participate in plant maintenance for Applied Wheat Pathology field experiments and phenotype wheat and perennial breeding material in the root rot field. In the lab, they will isolate and characterize the pathogens infecting plant root systems using morphological and molecular approaches. Lab 60%, Field 40% Mentor: Jessica Rupp Noller |
Using Viruses to Manage Plant Parasitic NematodesPlant parasitic nematodes are microscopic round worms that infect and damage plant roots. Plant Parasitic nematodes cost farmers billions of dollars in yield losses each year. Many farmers are forced to use costly and environmentally damaging pesticides to try and save their crops from these pests. but what if we could turn the tables on the nematodes by infecting them with a virus? The nematology lab here at Kansas State has identified viruses that are naturally present some plant parasitic nematode populations, and we need to figure out how they infect plant parasitic nematodes. In this 8-week research internship, a student will have the chance to conduct hands on experiments in the lab and greenhouse to try and figure out how viruses infect nematodes. The interns will have an opportunity to work independently alongside graduate students, postdocs, and technicians in the lab. They will learn how to collect and analyze a variety of different data types using molecular and bioinformatic tools. No prior experience will be necessary, only an interest in biology and a willingness to work with and learn from experienced scientists. Mentor: Will Rutter |
Engineering the Plant Epidermis for ResilienceThe outer cell layer, or epidermis of the plant, represents the first line of defense against the environment. The epidermis also plays a critical role in nutrient sensing, aligning growth with nutrient availability. Specific cell types, such as trichomes and giant cells, are determined by transcriptional regulators, but how are these regulators themselves controlled? Our lab studies protein-protein and protein-metabolite interactions involved in orchestrating growth in the Arabidopsis model organism. Interns will apply molecular techniques such as DNA assembly, gene editing, and gene expression analysis, to study specific interactions that are critical for epidermal traits. Interns will additionally learn skills for bioinformatic analysis of large data sets such as those from single-cell sequencing. Knowledge of the molecular mechanisms underlying epidermal cell-type differentiation will aid in engineering crops for resilience to environmental stresses in climate change scenarios. Mentor: Kathrin Schrick |
Ball buggy in the field
A formal ball buggy platform was designed after carefully considering various autonomous platform operational and robustness criteria. The prototype ball buggy is designed with wheels, vacuum system for cotton transfer and a bottom dropping-out system. An operational communication system was developed capable of communicating vehicle localization and operational data to a central hub located at the edge. The proposed plan is to further develop this system to interface with the central computer controlling the navigation and data acquisition. The data collected will be stored in packets with time stamps to communicate with the central hub. The initial data protocols will be utilized to send time-critical data (localization data) and vehicle operational data to the central hub. The data communication robustness will be evaluated by quantifying signal strength and the completeness of data received at the central hub. Mentor: Ajay Sharda |
Water for future crops
Data analytics, feedback between irrigation decisions and water resources sustainability, along with design and evaluation of decision support systems for improving crop water productivity with the coupling of anthropogenic and natural systems with applications in sustainable agriculture, water resources management, and applied hydrology. Interns will learn multiple data analysis and management techniques to download and get the data ready for input into process-based models and run the models to study the impact of the environment on water resource management for crop productivity. Mentor: Vaishali Sharda |
Improving insect pest control methods with molecular biologyThe Silver laboratory is interested in identifying new physiological mechanisms that can be exploited for insect pest control. Areas of exploration include physiological roles of extracellular vesicles as well as targets associated with the ability of blood feeding insects to find hosts or take blood meals. Interns will have the opportunity to learn molecular biology techniques, live cell imaging, cell culture, RNA interference, and/or insect behavioral assays while working alongside lab personnel and will develop independent research projects depending on their research interests. Lab100% Mentor: Kristopher Silver |
Computer Vision and Deep Learning for Pollinator MonitoringPollinators are active over large areas that can sometimes span miles. This makes understanding their foraging behavior challenging. Additionally, identifying many pollinators is difficult requiring collection and examination in the lab under a microscope. The process is time consuming, expensive, and requires lethal collection methods. Our lab is developing new techniques that automate this process by using computer vision to identify pollinators in the field. By combining computer vision algorithms with cameras on mobile devices and autonomous flying vehicles, we can crowdsource pollinator sightings and gather data over broad areas. Students will help us work on this project and develop a research project on pollinators that utilizes this technology. Mentor: Brian Spiesman |
Genetic mapping of natural variation in fungal pathogen traitsFusarium is a genus of filamentous fungi that contains many plant pathogens that can cause diseases on major crops and other agronomically important plants. Modern DNA sequencing technologies have brought the field of fungal genomics to even small research labs and have led to groundbreaking studies on the genetic changes that underlie natural trait variation. This project offers a number of opportunities to explore differences in distinct pathogen isolates. Participating students may conduct genetic crosses of isolates, conduct experiments to measure fungal isolate differences, such as how sensitive they are to fungicides, or help analyze data to associate genome-level genetic markers with important pathogen traits. This project has the potential to expand participants’ knowledge of: bioinformatics such as the processing of next-generation sequence data and the genomic mapping of trait differences; molecular techniques such as nucleic acid extraction and DNA sequencing; and microbiological techniques such as culturing and crossing fungal isolates and measuring growth and spore production rates. Mentor: Chris Toomajian |
Super plants...The advanced modern biotechnology changed many aspects in agriculture. Recent rapidly developed genome editing technology such CRISPR/Cas9 is changing the world in agriculture as well as human medicine. In this project, you will learn how to design, construct, and use engineered plasmid vectors to introduce new traits by expressing genes and/or genome editing in plants (wheat or soybean). You will gain knowledge about the interaction between viral pathogen and plant host. You will gain skills in molecular biology, plant transformation and micropropagation, as well as DNA/RNA extraction, and virus biology, virus detection and analysis. Lab 80%, Greenhouse 20% Mentor: Harold Trick |
Dissecting plant stressAn REEU student will work in the model plant Arabidopsis thaliana to characterize the function of an uncharacterized gene involved in response to abiotic plant stress. The gene will be chosen from the genes identified in mutant screens and genome wide association studies designed to detect genes associated with lipid changes in response to wounding, cold, or freezing. The functional characterization will include analysis of multiple mutants in the gene of interest in comparison with wild-type plants. The student will confirm the presence of the mutation, measure gene expression in wild-type and mutant plants, analyze lipid changes under basal conditions and in the presence of environmental stressors, and measure relevant physiological and visible phenotypes. The work will shed light on mechanisms of abiotic stress, while providing the student with a solid foundation in basic plant science research. Mentor: Ruth Welti |
Contact Us
Project Director
Dr. Megan Kennelly![]()
Project Coordinator
Dr. Myron Bruce![]()
Project Co-Director
Dr. David Cook![]()
 Have you heard of DNA sequencing but you’re not quite sure what it is or how it is used? Maybe you’ve had a class or heard a news article that discussed genomics or a genome, but it is not clear what that means or how it was created. Complete an internship in the Cook lab as part of the K- State REEU program and get hands on experience extracting fungal DNA, preparing sequencing libraries for long-read sequencing, and learn basic bioinformatic skills to analyze the data. You do not need prior experience, and we will work with you to gain proficiency and knowledge about DNA sequencing and the types of experimental questions that can be addressed.
Have you heard of DNA sequencing but you’re not quite sure what it is or how it is used? Maybe you’ve had a class or heard a news article that discussed genomics or a genome, but it is not clear what that means or how it was created. Complete an internship in the Cook lab as part of the K- State REEU program and get hands on experience extracting fungal DNA, preparing sequencing libraries for long-read sequencing, and learn basic bioinformatic skills to analyze the data. You do not need prior experience, and we will work with you to gain proficiency and knowledge about DNA sequencing and the types of experimental questions that can be addressed.  Just like we all have some interesting relatives, wheat has some wild in-laws of its own! Bread wheat was domesticated in Central Asia around Iran and Iraq and the wild relatives can still be found there today. Though they don’t really even look like wheat and aren’t any good for making bread, the wild relatives have important genes for disease resistance. Using chromosome engineering and bioinformatics, we are transferring these useful genes into wheat to make stronger wheat plants that are resistant to disease and other stress.
Just like we all have some interesting relatives, wheat has some wild in-laws of its own! Bread wheat was domesticated in Central Asia around Iran and Iraq and the wild relatives can still be found there today. Though they don’t really even look like wheat and aren’t any good for making bread, the wild relatives have important genes for disease resistance. Using chromosome engineering and bioinformatics, we are transferring these useful genes into wheat to make stronger wheat plants that are resistant to disease and other stress.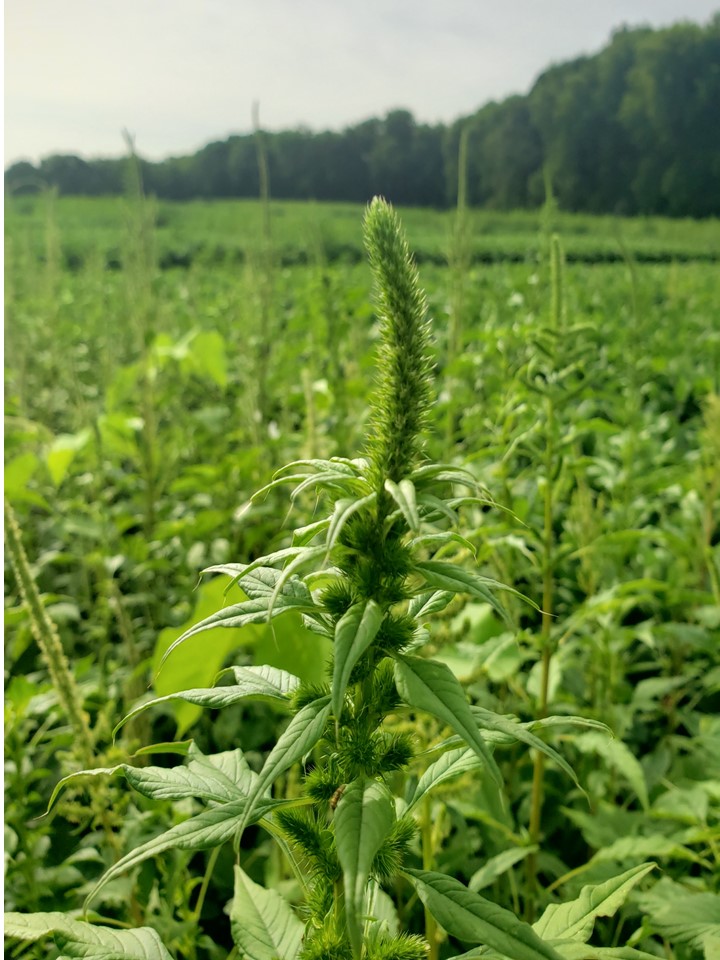 Weeds are one of the greatest challenges farmers have. Weed scientists work to find effective and economical solutions to farmers’ weed management challenges and share that information with farmers and agribusinesses. In this internship, students will have the opportunity to work in the field on projects related to herbicide evaluation as well as nonchemical weed management practices. In addition, students will have the opportunity to participate in Extension-related activities, including summarizing data. If you enjoy biology or chemistry, gathering information, working outdoors, and helping people, this internship might be for you!
Weeds are one of the greatest challenges farmers have. Weed scientists work to find effective and economical solutions to farmers’ weed management challenges and share that information with farmers and agribusinesses. In this internship, students will have the opportunity to work in the field on projects related to herbicide evaluation as well as nonchemical weed management practices. In addition, students will have the opportunity to participate in Extension-related activities, including summarizing data. If you enjoy biology or chemistry, gathering information, working outdoors, and helping people, this internship might be for you!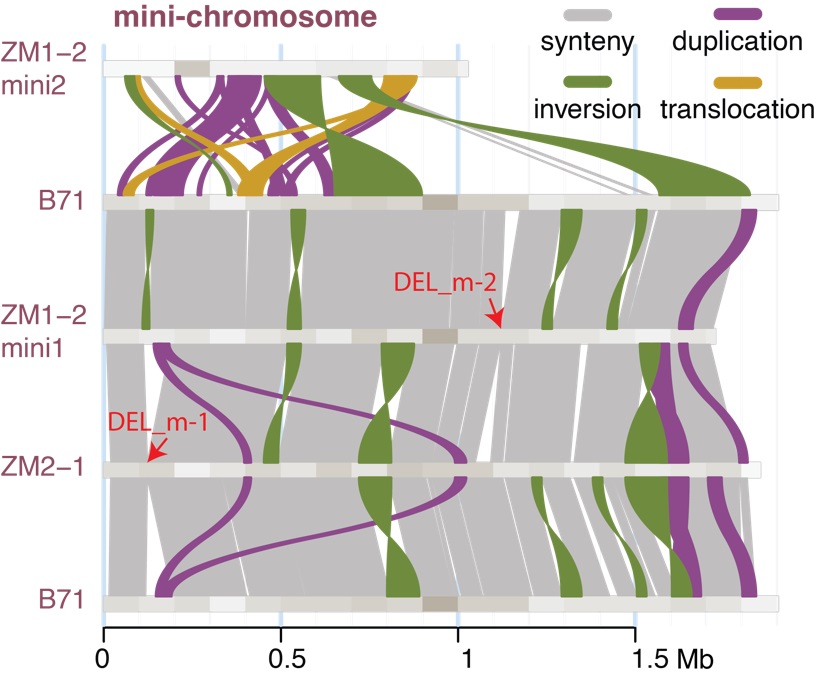 Genome sequencing technology has become an important tool for understanding evolution, genetic bases
Genome sequencing technology has become an important tool for understanding evolution, genetic bases  Park team focuses on the development of new tools to disrupt insect endocrine system based on the knowledge of hormone receptors revealed in comparative genomics. Current project includes high throughput screening of chemical compounds that act on insect specific ecdysis triggering hormone receptor and mite specific neuropeptide receptors. Students will be trained for molecular biology and data analyses of bioassay depending on the background and the interests of the students assigned. Most of all, the students will have opportunities to play with arthropods, learn the fun biology, and perform modern molecular techniques to solve the problems in pest control.
Park team focuses on the development of new tools to disrupt insect endocrine system based on the knowledge of hormone receptors revealed in comparative genomics. Current project includes high throughput screening of chemical compounds that act on insect specific ecdysis triggering hormone receptor and mite specific neuropeptide receptors. Students will be trained for molecular biology and data analyses of bioassay depending on the background and the interests of the students assigned. Most of all, the students will have opportunities to play with arthropods, learn the fun biology, and perform modern molecular techniques to solve the problems in pest control.